Abstract
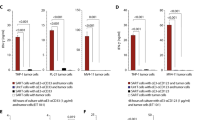
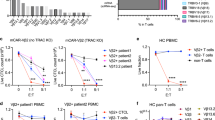
Solid organ transplant (SOT) recipients receive immunosuppressive drugs (ISDs) and are susceptible to developing severe COVID-19. Here, we analyze the Spike-specific T-cell response after 3 doses of mRNA vaccine in a group of SOT patients (n = 136) treated with different ISDs. We demonstrate that a combination of a calcineurin inhibitor (CNI), mycophenolate mofetil (MMF), and prednisone (Pred) treatment regimen strongly suppressed the mRNA vaccine-induced Spike-specific cellular response. Such defects have clinical consequences because the magnitude of vaccine-induced Spike-specific T cells was directly proportional to the ability of SOT patients to rapidly clear SARS-CoV-2 after breakthrough infection. To then compensate for the T-cell defects induced by immunosuppressive treatment and to develop an alternative therapeutic strategy for SOT patients, we describe production of 6 distinct SARS-CoV-2 epitope-specific ISD-resistant T-cell receptor (TCR)-T cells engineered using the mRNA electroporation method with reactivity minimally affected by mutations occurring in Beta, Delta, Gamma, and Omicron variants. This strategy with transient expression characteristics marks an improvement in the immunotherapeutic field and provides an attractive and novel therapeutic possibility for immunosuppressed COVID-19 patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
For original data, please contact Antonio Bertoletti (antonio@duke-nus.edu.sg).
References
Pan Q, Tilanus HW, Metselaar HJ, Janssen HL, van der Laan LJ. Virus-drug interactions-molecular insight into immunosuppression and HCV. Nat Rev Gastroenterol Hepatol. 2012;9:355–62.
Raja MA, Mendoza MA, Villavicencio A, Anjan S, Reynolds JM, Kittipibul V, et al. COVID-19 in solid organ transplant recipients: A systematic review and meta-analysis of current literature. Transpl Rev. 2021;35:100588.
Pereira MR, Mohan S, Cohen DJ, Husain SA, Dube GK, Ratner LE, et al. COVID-19 in solid organ transplant recipients: Initial report from the US epicenter. Am J Transplant. 2020;20:1800–8.
Simioli F, Martino M, Annunziata A, Carannante N, Fiorentino G. Therapeutic approach for severe COVID-19 and immunocompromised patients. A case series. Respir Med Case Rep. 2021;33:101397.
Avery RK. Update on COVID-19 therapeutics for solid organ transplant recipients, including the omicron surge. Transplantation. 2022;106:1528–37.
van Laarhoven A, Kurver L, Overheul GJ, Kooistra EJ, Abdo WF, van Crevel R, et al. Interferon gamma immunotherapy in five critically ill COVID-19 patients with impaired cellular immunity: a case series. Medicne. 2021;2:1163–70.e2.
Bertrand D, Hamzaoui M, Lemée V, Lamulle J, Hanoy M, Laurent C, et al. Antibody and T cell response to SARS-CoV-2 messenger RNA BNT162b2 vaccine in kidney transplant recipients and hemodialysis patients. J Am Soc Nephrol. 2021;32:2147–52.
Rincon-Arevalo H, Choi M, Stefanski AL, Halleck F, Weber U, Szelinski F, et al. Impaired humoral immunity to SARS-CoV-2 BNT162b2 vaccine in kidney transplant recipients and dialysis patients. Sci Immunol. 2021;6:eabj1031.
Gao Y, Cai C, Wullimann D, Niessl J, Rivera-Ballesteros O, Chen P, et al. Immunodeficiency syndromes differentially impact the functional profile of SARS-CoV-2-specific T cells elicited by mRNA vaccination. Immunity. 2022;55:1732–46.e5.
Baang JH, Smith C, Mirabelli C, Valesano AL, Manthei DM, Bachman MA, et al. Prolonged severe acute respiratory syndrome coronavirus 2 replication in an immunocompromised patient. J Infect Dis. 2021;223:23–27.
Helleberg M, Niemann CU, Moestrup KS, Kirk O, Lebech AM, Lane C, et al. Persistent COVID-19 in an immunocompromised patient temporarily responsive to two courses of remdesivir therapy. J Infect Dis. 2020;222:1103–7.
Nakajima Y, Ogai A, Furukawa K, Arai R, Anan R, Nakano Y, et al. Prolonged viral shedding of SARS-CoV-2 in an immunocompromised patient. J Infect Chemother. 2021;27:387–9.
Cajanding R. Immunosuppression following organ transplantation. Part 1: mechanisms and immunosuppressive agents. Br J Nurs. 2018;27:920–7.
Casadevall A, Focosi D. SARS-CoV-2 variants resistant to monoclonal antibodies in immunocompromised patients constitute a public health concern. J Clin Investig. 2023;133:e168603.
Tan AT, Linster M, Tan CW, Le Bert N, Chia WN, Kunasegaran K, et al. Early induction of functional SARS-CoV-2-specific T cells associates with rapid viral clearance and mild disease in COVID-19 patients. Cell Rep. 2021;34:108728.
Feuchtinger T, Opherk K, Bethge WA, Topp MS, Schuster FR, Weissinger EM, et al. Adoptive transfer of pp65-specific T cells for the treatment of chemorefractory cytomegalovirus disease or reactivation after haploidentical and matched unrelated stem cell transplantation. Blood. 2010;116:4360–7.
Comoli P, Labirio M, Basso S, Baldanti F, Grossi P, Furione M, et al. Infusion of autologous Epstein-Barr virus (EBV)-specific cytotoxic T cells for prevention of EBV-related lymphoproliferative disorder in solid organ transplant recipients with evidence of active virus replication. Blood. 2002;99:2592–8.
Martits-Chalangari K, Spak CW, Askar M, Killian A, Fisher TL, Atillasoy E, et al. ALVR109, an off-the-shelf partially HLA matched SARS-CoV-2-specific T cell therapy, to treat refractory severe COVID-19 pneumonia in a heart transplant patient: case report. Am J Transplant. 2022;22:1261–5.
Papadopoulou A, Karavalakis G, Papadopoulou E, Xochelli A, Bousiou Z, Vogiatzoglou A, et al. SARS-CoV-2-specific T cell therapy for severe COVID-19: a randomized phase 1/2 trial. Nat Med. 2023;29:2019–29.
Vasileiou S, Hill L, Kuvalekar M, Workineh AG, Watanabe A, Velazquez Y, et al. Allogeneic off-the-shelf SARS-CoV-2-specific T cells (ALVR109) for the treatment of COVID-19 in high-risk patients. Haematologica. 2023;108:1840–50.
Peter L, Wendering DJ, Schlickeiser S, Hoffmann H, Noster R, Wagner DL, et al. Tacrolimus-resistant SARS-CoV-2-specific T cell products to prevent and treat severe COVID-19 in immunosuppressed patients. Mol Ther Methods Clin Dev. 2022;25:52–73.
Hafezi M, Lin M, Chia A, Chua A, Ho ZZ, Fam R, et al. Immunosuppressive drug-resistant armored T-cell receptor T cells for immune therapy of HCC in liver transplant patients. Hepatology. 2021;74:200–13.
De Angelis B, Dotti G, Quintarelli C, Huye LE, Zhang L, Zhang M, et al. Generation of Epstein-Barr virus-specific cytotoxic T lymphocytes resistant to the immunosuppressive drug tacrolimus (FK506). Blood. 2009;114:4784–91.
Yam P, Jensen M, Akkina R, Anderson J, Villacres MC, Wu J, et al. Ex vivo selection and expansion of cells based on expression of a mutated inosine monophosphate dehydrogenase 2 after HIV vector transduction: effects on lymphocytes, monocytes, and CD34+ stem cells. Mol Ther. 2006;14:236–44.
Koh S, Shimasaki N, Suwanarusk R, Ho ZZ, Chia A, Banu N, et al. A practical approach to immunotherapy of hepatocellular carcinoma using T cells redirected against hepatitis B virus. Mol Ther Nucleic Acids. 2013;2:e114.
Bertoletti A, Tan AT. Challenges of CAR- and TCR-T cell-based therapy for chronic infections. J Exp Med. 2020;217:e20191663.
Kah J, Koh S, Volz T, Ceccarello E, Allweiss L, Lütgehetmann M, et al. Lymphocytes transiently expressing virus-specific T cell receptors reduce hepatitis B virus infection. J Clin Investig. 2017;127:3177–88.
Polidoro RB, Hagan RS, de Santis Santiago R, Schmidt NW. Overview: systemic inflammatory response derived from lung injury caused by SARS-CoV-2 infection explains severe outcomes in COVID-19. Front Immunol. 2020;11:1626.
Lim JME, Hang SK, Hariharaputran S, Chia A, Tan N, Lee ES, et al. A comparative characterization of SARS-CoV-2-specific T cells induced by mRNA or inactive virus COVID-19 vaccines. Cell Rep. Med. 2022;3:100793.
Peng Y, Felce SL, Dong D, Penkava F, Mentzer AJ, Yao X, et al. An immunodominant NP105-113-B*07:02 cytotoxic T cell response controls viral replication and is associated with less severe COVID-19 disease. Nat Immunol. 2022;23:50–61.
Goh YS, Chavatte JM, Lim Jieling A, Lee B, Hor PX, Amrun SN, et al. Sensitive detection of total anti-Spike antibodies and isotype switching in asymptomatic and symptomatic individuals with COVID-19. Cell Rep. Med. 2021;2:100193.
Goh YS, Rouers A, Fong SW, Zhuo NZ, Hor PX, Loh CY, et al. Waning of specific antibodies against Delta and Omicron variants five months after a third dose of BNT162b2 SARS-CoV-2 vaccine in elderly individuals. Front Immunol. 2022;13:1031852.
Goh YS, Ng LFP, Renia L. A flow cytometry-based assay for serological detection of anti-spike antibodies in COVID-19 patients. STAR Protoc. 2021;2:100671.
Koutsakos M, Reynaldi A, Lee WS, Nguyen J, Amarasena T, Taiaroa G, et al. SARS-CoV-2 breakthrough infection induces rapid memory and de novo T cell response. Immunity. 2023;56:879–892.e4.
Ricciardelli I, Brewin J, Lugthart G, Albon SJ, Pule M, Amrolia PJ. Rapid generation of EBV-specific cytotoxic T lymphocytes resistant to calcineurin inhibitors for adoptive immunotherapy. Am J Transplant. 2013;13:3244–52.
Tan AT, Yang N, Lee Krishnamoorthy T, Oei V, Chua A, Zhao X, et al. Use of expression profiles of HBV-DNA integrated into genomes of hepatocellular carcinoma cells to select T cells for immunotherapy. Gastroenterology. 2019;156:1862–76.e9.
Qasim W, Brunetto M, Gehring AJ, Xue SA, Schurich A, Khakpoor A, et al. Immunotherapy of HCC metastases with autologous T cell receptor redirected T cells, targeting HBsAg in a liver transplant patient. J Hepatol. 2015;62:486–91.
Tometten I, Landmann S, Kantauskaite M, Lamberti J, Hillebrandt J, Müller L, et al. Factors associated with vaccine-induced T cell immune responses against SARS-CoV-2 in kidney transplant recipients. J Infect Dis. 2022;227:641–50.
Scurr MJ, Lippiatt G, Capitani L, Bentley K, Lauder SN, Smart K, et al. Magnitude of venous or capillary blood-derived SARS-CoV-2-specific T cell response determines COVID-19 immunity. Nat Commun. 2022;13:5422.
Choi B, Choudhary MC, Regan J, Sparks JA, Padera RF, Qiu X, et al. Persistence and evolution of SARS-CoV-2 in an immunocompromised host. N Engl J Med. 2020;383:2291–3.
Tan AT, Lim JM, Le Bert N, Kunasegaran K, Chia A, Qui MD, et al. Rapid measurement of SARS-CoV-2 spike T cells in whole blood from vaccinated and naturally infected individuals. J Clin Investig. 2021;131:e152379.
Liu J, Yu J, McMahan K, Jacob-Dolan C, He X, Giffin V, et al. CD8 T cells contribute to vaccine protection against SARS-CoV-2 in macaques. Sci Immunol. 2022;7:eabq7647.
Basar R, Uprety N, Ensley E, Daher M, Klein K, Martinez F, et al. Generation of glucocorticoid-resistant SARS-CoV-2 T cells for adoptive cell therapy. Cell Rep. 2021;36:109432.
Canas CA. The triggering of post-COVID-19 autoimmunity phenomena could be associated with both transient immunosuppression and an inappropriate form of immune reconstitution in susceptible individuals. Med Hypotheses. 2020;145:110345.
Sharma C, Bayry J. High risk of autoimmune diseases after COVID-19. Nat Rev Rheumatol. 2023;19:399–400.
Quach D, Lulla P, Rooney CM. Banking on virus-specific T-cells (VSTS) to fulfill the need for “off the shelf” cell therapies. Blood. 2022;139:799–802.
Bertoletti A, Le Bert N, Tan AT. SARS-CoV-2-specific T cells in the changing landscape of the COVID-19 pandemic. Immunity. 2022;55:1764–78.
Acknowledgements
We would like to acknowledge the contribution of the Singapore National University Centre for Organ Transplantation team members who helped recruit patients: AV, AL, and WKK. We thank all voluntary blood donors for their donations. We would like to thank the members of AB’s lab for their insights and critique. Finally, we would also like to thank Dr. Yongxu Lu and Prof. Geoffrey L. Smith from the Department of Pathology, University of Cambridge, U.K., for supplying the vaccinia virus-expressing Spike and Nucleocapsid proteins. This study was supported by research funding from the Singapore Ministry of Health’s National Medical Research Council MOH-000019 (MOH-StaR17Nov-001) to Antonio Bertoletti. Part of this work was also supported by the A*ccelerate GAP-funded project (ACCL/19-GAP064-R20H-H) from the Agency of Science, Technology and Research (A*STAR), the Singapore National Medical Research Council COVID-19 Research Fund (COVID19RF-011) and a Start-up University Grant from the Ministry of Education (Singapore) to Laurent Renia. YSG was supported by a Career Development Fund award by A*STAR (SC35/22-805100).
Author information
Authors and Affiliations
Contributions
QC and AB conceptualized and designed the experiments. QC, AC, SKH, KK, YC.P, FG, and JLC performed the experiments. AL, WKK, and AV recruited the transplant patients and collected the samples. ZH and LEW performed the single-cell TCR sequencing experiment. YSG, CYL, and LR generated constructs for EBV-B cells expressing the ancestral and the Omicron variant spike proteins. QC, YCP, ATT, TD, NLB, and AB analyzed and interpreted the data. QC prepared the figures. QC and AB wrote the manuscript. AB designed and coordinated the study and provided funding.
Corresponding author
Ethics declarations
Competing interests
AB is a cofounder of and ATT consults for Lion TCR, a biotech company developing T-cell receptors for treatment of virus-related diseases and cancers. ZH and LEW are employees of Lion TCR Pte. Ltd. None of the other authors has any competing interests related to the study.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, Q., Chia, A., Hang, S.K. et al. Engineering immunosuppressive drug-resistant armored (IDRA) SARS-CoV-2 T cells for cell therapy. Cell Mol Immunol 20, 1300–1312 (2023). https://doi.org/10.1038/s41423-023-01080-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41423-023-01080-3