Abstract
Glycyrrhiza, a genus of perennial medicinal herbs, has been traditionally used to treat human diseases, including respiratory disorders. Functional analysis of genes involved in the synthesis, accumulation, and degradation of bioactive compounds in these medicinal plants requires accurate measurement of their expression profiles. Reverse transcription quantitative real-time PCR (RT-qPCR) is a primary tool, which requires stably expressed reference genes to serve as the internal references to normalize the target gene expression. In this study, the stability of 14 candidate reference genes from the two congeneric species G. uralensis and G. inflata, including ACT, CAC, CYP, DNAJ, DREB, EF1, RAN, TIF1, TUB, UBC2, ABCC2, COPS3, CS, R3HDM2, were evaluated across different tissues and throughout various developmental stages. More importantly, we investigated the impact of interactions between tissue and developmental stage on the performance of candidate reference genes. Four algorithms, including geNorm, NormFinder, BestKeeper, and Delta Ct, were used to analyze the expression stability and RefFinder, a comprehensive software, provided the final recommendation. Based on previous research and our preliminary data, we hypothesized that internal references for spatio-temporal gene expression are different from the reference genes suited for individual factors. In G. uralensis, the top three most stable reference genes across different tissues were R3HDM2, CAC and TUB, while CAC, CYP and ABCC2 were most suited for different developmental stages. CAC is the only candidate recommended for both biotic factors, which is reflected in the stability ranking for the spatio (tissue)-temporal (developmental stage) interactions (CAC, R3HDM2 and DNAJ). Similarly, in G. inflata, COPS3, R3HDM2 and DREB were selected for tissues, while RAN, COPS3 and CS were recommended for developmental stages. For the tissue-developmental stage interactions, COPS3, DREB and ABCC2 were the most suited reference genes. In both species, only one of the top three candidates was shared between the individual factors and their interactions, specifically, CAC in G. uralensis and COPS3 in G. inflata, which supports our overarching hypothesis. In summary, spatio-temporal selection of reference genes not only lays the foundation for functional genomics research in Glycyrrhiza, but also facilitates these traditional medicinal herbs to reach/maximize their pharmaceutical potential.
Similar content being viewed by others
Introduction
Licorice or liquorice is the common name of Glycyrriza uralensis Fischer, Glycyrrhiza glabra Linné or Glycyrrhiza inflata Batalin, which are herbaceous perennial plants of the bean family Fabaceae native to the western Asia and southern Europe1. Besides its ecological values for windbreak and sand fixation, both roots and shoots of licorice compose specialized bioactive compounds/molecules with pharmaceutical potential. Licorice root extracts have been used in herbalism and traditional medicine and presented anti-carcinogenic2,3, anti-inflammatory, anti-fungal, anti-piroplasmic and cytotoxic activities4. More recently, glycyrrhizin, the most important bioactive triterpenoid saponin in licorice roots, is under the consideration for treating COVID-19 infection caused respiratory syndrome5. And licorice shoots are a kind of high-quality forage grass because of their high content of coarse fiber and flavonoid6. Their beneficial effects on human health has made licorice a valuable trade item. However, these bioactive compounds in licorice with pharmaceutical interest, such as glycyrrhizin or flavonoid, are typically in minute quantities7. Therefore, a better understanding of pathways associated with biosynthesis, regulation, and accumulation of these phytochemicals becomes a key step to reach the pharmaceutical potential of licorice8,9,10.
Spatio-temporal gene expression is the activation of genes within specific tissues of an organism at specific times during development (WIKIPEDIA). Many key genes only express in certain tissues and at certain developmental stages in response to both internal and external cues to ensure the accomplishment of each step in plant life cycles11,12. In addition, accumulation of valuable bioactive constituents in many functional plants is tissue-specific and meanwhile, only happens at specific developmental stages. One well-known example are ginsenosides in ginseng, which accumulate specifically in roots and rhizomes in “mature” plants, while little could be detected at juvenile stage in these perennial plants13. Consistently, the expression of regulatory genes and biosynthetic genes of ginsenosides is also spatio-temporal specific14. A growing number of studies have been conducted to screen genes involved in the same metabolic pathways by co-expression network15,16. Spatio-temporal gene expression profiling may provide important clues for future functional analyses of genes in non-model organisms.
Reverse transcription quantitative real-time PCR (RT-qPCR) is a method for accurate expression analysis and comparisons of small numbers of genes among various experimental samples17,18. Because of the accuracy and sensitivity of RT-qPCR suitable internal references for data normalization in RT-qPCR analysis are prerequisite to obtain reliable results19. Expression of suitable reference genes should be constant under the experimental conditions to be tested in a specific research19. Housekeeping genes, due to their stability and indispensable function for survival, are the typical first choice for reference gene selection20,21, and stably expressed genes in RNA-seq experiments might also be good candidates. That being said, no “universal” reference gene has been verified to be stably expressed across all given experimental conditions22,23. Consequently, selection of appropriate reference genes is required for a standardized RT-qPCR procedure following the MIQE (Minimum Information for publication of Quantitative real time PCR Experiments) guidelines24.
In licorice, the types and contents of many bioactive compounds varied remarkably among different tissues at different developmental stages. Glycyrrhizin, the most important bioactive triterpenoid saponin in licorice roots, are predominantly accumulated in roots and rhizomes25, and accumulated to higher levels in summer than in winter26. Spatio-temporal spcific expression of related biosynthetic and regulatory genes should be the cause of these spatial-temporol accumulation of bioactive compounds. For example, the key genes in glycyrrhizin biosynthetic pathway, β-amyrin synthase (β-AS), β-amyrin 11-oxidase (CYP88D6), 11-oxo-β-amyrin 30-oxidase (CYP72A154), are mainly expressed in roots and rhizomes, whereas no transcripts were observed in leaves8,9,27. Therefore, the screening of reference genes across different tissues or throughout different developmental stages, more importantly, the spatio (tissue)-temporal (developmental stage) interactions is of great importance for the study of gene functions in licorice.
Selection of reference genes, as of now, has been focused on a single factor/dimension, i.e., time (developmental stage) or space (tissue). However, the expression of genes is constantly under the influences of multiple factors/dimensions. Here, to better understand the spatio-temporal gene expression patterns in licorice, we investigated the expression profiles of 14 candidate reference genes in this study. Although Maroufi28 previously studied reference genes under the drought stresses in G. glabra, information concerning suitable reference genes is still lacking in licorice, especially for congeneric species, G. uralensis or G. inflata.
Based on previous research and our preliminary data, we hypothesized that internal references for spatio-temporal gene expression are different from the reference genes suited for individual conditions. To examine this hypothesis, we 1) evaluated the stability of 14 candidate genes, which were stably expressed housekeeping genes derived from our RNA-seq analysis; 2) selected optimal reference genes under the conditions of different tissues, developmental stages, and tissue × developmental stages, respectively; 3) compared the suitable reference genes under different experimental conditions and between the two congenic Glycyrrhiza species, and finally, we summarized the reference genes previously used within Leguminosae plants.
Materials and methods
Plant materials and growth conditions
Two-year-old licorice plants (G. uralensis and G. inflata) were grown in the test field at the Northwest Biological Agricultural Center, Chinese Academy of Sciences (Ningxia, China). Roots, rhizomes and leaves were collected in April (returning green), May (rapid growth and flowering), July (seed setting), and October (aging) (Fig. 1). All samples were flash frozen in liquid nitrogen, shipped to Guangzhou in dry ice and stored at -80 ℃ for RNA extraction. All experiments were carried out with three biological replicates.
The morphology of different tissues under different developmental stages of G. uralensis and G. inflata. (A) schematic drawings of G. uralensis and G. inflate, respectively; (B) morphology of root and rhizome of G. urelensis and G. inflata, respectively; (C) G. urelensis and G. inflate, respectively, in different developmental stages, including Returning green stage (April), Rapid growth and flowering stage (May), Seed setting (July), and Senescence stage (October).
RNA isolation and cDNA synthesis
Total RNA was isolated using HiPure Total RNA Mini Kit (Code No. R4151-03, Magen, China) according to the manufacturer’s instruction. The RNA quality and concentration were measured with agarose gel electrophoresis and spectrophotometer (NanoDrop 2000, Thermo, USA). Removal of genomic DNA contamination and first strand cDNA synthesis were performed using PrimeScript RT reagent Kit with gDNA Eraser (Code No. RR047A, Takara, Dalian, China).
Candidate reference gene screening and primer design
A total of 14 candidate reference genes was included in this study, in which ten, (Actin1 (ACT), Clathrin complex AP1 (CAC), Cyclophilin (CYP), Heat-shock protein 40 (DNAJ), Dehydration responsive element binding gene (DREB), Translation elongation factor1 (EF1), Ras related protein (RAN), Translation initiation factor (TIF1), β-Tubulin (TUB), Ubiquitin-conjugating enzyme E2 (UBC2),) were selected in previous studies28,29, and the other four (ATP binding-box transpoter 2 (ABCC2), COP9 signal compex subunit 3 (COPS3), Citrate synthase (CS), R3H domain protein 2 (R3HDM2) were selected from a RNA-seq dataset (SRA accession: PRJNA574093). RT-qPCR primers were designed using PrimerQuest Tool, INTEGRATED DNA TECHNOLOGIES (IDT) (https://sg.idtdna.com/Primerquest/Home/Index) based on the following parameters: Melting temperature (Tm) of 59–65 ℃; GC content of 45–55%; optimum length of 17–30 bp and amplicon length of 50–200 bp. All primers were synthesized by TSINGKE Company (Guangzhou, China). Detailed information listed in Table 1.
RT-qPCR analysis
The RT-qPCR were carried out in 384-well blocks using TB Green Premix Ex Taq II (Tli RNaseH Plus) (Code No. RR820D, Takara, Dalian, China) on LightCycler 480 (Roche, Switzerland) according to manufacturers’ instructions. Three technical repeats were carried out for each sample.
Expression stability analysis of candidate reference genes
The slope of the standard curve of a cDNA tenfold dilution series was constructed to calculate the PCR amplification efficiency (E), and the E value was obtained according to the equation E = [10(−1/slope) − 1] × 100. Expression stability of the 14 candidate reference genes were evaluated by four Microsoft Excel-based computational programs, geNorm30, NormFinder31, BestKeeper32 and Delta CT33. geNorm method ranks the expression stability by M value for each reference gene, and the smaller the M value, the more stable the gene. Based on NormFinder, the gene expression stability was calculated by the SV value. BestKeeper calculates the stability of candidate genes by the Pearson correlation coefficient (r), while the stability of the genes is evaluated by the pair-wise comparisons.
Selection of optimal reference genes under different experimental conditions
The RefFinder, a comprehensive system to integrate the currently available major computational programs (geNorm, Normfinder, BestKeeper, and Delta Ct method) to compare and rank the stability of candidate reference genes, was used for the overall ranking of the candidate reference genes34. Based on the rankings from the Microsoft Excel-based computational programs, RefFinder assigns an appropriate weight to an individual gene and calculates the geometric mean of their weights for the overall final ranking.
Comparison of the suitable reference genes under different experimental conditions and between the two congenic Glycyrrhiza species
The top three most suitable reference genes selected by RefFinder under the conditions of different tissues, developmental stages, and tissue × developmental stages were compared; and the suitable reference genes under the same experimental condition between the two congenic Glycyrrhiza species were also analyzed. The results were visualized by Venn Diagrams, and it was plotted using the OmicShare tools, a free online platform for data analysis (www.omicshare.com/tools).
Validation of recommended reference genes
To confirm the suitability of the reference genes recommended in the present study, we measured the differential expression of a specific licorice gene with a known expression profile under different tissues. Licorice β-amyrin synthase (β-AS, GenBank Accession Number: FJ627179), is a key gene in glycyrrhizin biosynthesis and mainly expressed in root organs8, 27, we thus chose to validate the reliability of the selected reference genes in different treatment conditions. The primers of β-AS used for RT-qPCR were listed in Table 1. The expression level of β-AS was analyzed by seven normalization ways, including the most stable, the top two most stable, the top three most stable, the least stable, the top two least stable, the top three least stable, and all the candidate reference genes. One way ANOVA was carried out to evaluate the expression level of β-AS under different normalization conditions (SPSS statistics 22.0 software, IBM, United States).
Survey of the reference genes used within Leguminosae plants
Reference genes reported previously in Leguminosae species was selected by searching “NCBI” or “Web of Science” with the key words “Leguminosae” and “reference gene”. The top three most suitable reference gene was considered to calculate the recommendation frequency for developmental stages and tissues, respectively, among Leguminosae species.
Results
Primer specificity and RT-qPCR amplification efficiency
The specificity of the primers for all 14 candidate reference genes (ACT, CAC, CYP, DNAJ, DREB, EF1, RAN, TIF1, TUB, UBC2, ABCC2, COPS3, CS, R3HDM2) were determined by a single PCR product of expected size, and further confirmed by a single peak in the melting-curve analysis. The efficiency of the PCR amplification (E) was calculated from the standard curve by making a dilution series with mixed samples. The E value of the reported reference genes ranged from 67.28% (CAC) to 116.84% (CYP) and correlation coefficients varied from 0.9526 (CAC) to 0.9976 (CYP) (Table 1). Three of them, CAC, TIF1 and TUB, presented unmatched E values which did not meet the requirements (90%—110%) for RT-qPCR analysis. The E value of the selected candidate reference genes from RNA-Seq data ranged from 90.63% (CAC) to 107.58% (CYP) and correlation coefficients varied from 0.9887 (CS) to 0.9963 (COPS3) (Table 1), and all these primers are suitable for RT-qPCR analysis32.
Expression profiling of candidate reference genes
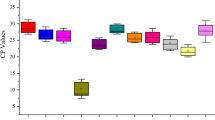
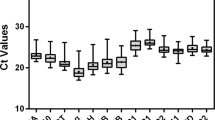
The expression levels of the candidate reference genes were determined as quantification cycle (Cq) values, and the transcripts of these genes showed different levels of abundance in G. uralensis and G. inflata (Tables S1 and S2). The mean Cq values of the genes ranged from 20—27, with the majority lying between 23 and 26 across all tested samples in G. uralensis, and the mean Cq values of the genes ranged from 19—25, with the majority lying between 22 and 25 across all tested samples in G. inflata (Fig. 2, Tables S3 and S4). So the expression level of these candidate genes were much higher in tested samples in G. inflata than in G. uralensis, and they were more stable in G. inflata than in G. uralensis.
Distribution of threshold cycle (Cq) values for candidate reference genes. Distribution of threshold cycle (Cq) values of G. uralensis and G. inflata in all samples. Boxes: the interquartile range; lines across the boxes: median; lines above and below the boxes: the maximum and minimum values; black dots: outliers.
TIF1 had the lowest Cq both in G. uralensis (mean Ct of 20.30) and G. inflata (mean Cq of 19.94), indicating the highest expression level of TIF1 in the two species, while EF1 (mean Cq of 27.50) or DREB (mean Cq of 25.56) was expressed at low levels in G. uralensis or G. inflata, respectively (Tables S3 and S4).
CAC showed the least gene expression variation both in G. uralensis (coefficient of variation (CV) of 6.74%) and G. inflata (CV of 2.92%), while surprisingly, a commonly used reference gene, ACT, was the most variable across all samples (CV of 10.45%) in G. uralensis (Table S3), and TIF1 (9.60%) was the most variable across all samples in G. inflata (Table S4).
Expression stability of candidate reference genes
The expression profiles of the 14 candidate reference genes in G. uralensis and G. inflata roots, rhizomes, and leaves across all experiments in this study were analyzed using geNorm, NormFinder, BestKeeper, Delta CT, and RefFinder (Tables 2 and 3).
For different developmental stages, four key growth periods of licorice were selected and measured, including Returning green stage in April; Rapid growth and flowering stage in May; Seed setting stage in July; and Senescence stage in October (Fig. 1). In G. uralensis, CAC, CYP, COPS3 were the most stable reference genes recommended by geNorm, whereas CAC, ABCC2, CYP by NormFinder, DNAJ, COPS3, CAC by the BestKeeper, and CAC, ABCC2, DREB by Delta CT (Table 2). In G. inflata, the top three most stable candidate reference genes were RAN, COPS3, CS identified by all four methods, geNorm, NormFinder, BestKeeper and Delta CT (Table 3).
For different tissues, three tissues (the roots, rhizomes and leaves) were tested (Fig. 1). In G. uralensis, the top three most stable reference genes recommended by geNorm and NormFinder were CAC, R3HDM2 and TUB, while COPS3, RAN and TIF1 by BestKeeper, R3HDM2, TUB and CAC by Delta CT (Table 2). In G. inflata, DREB, R3HDM2 and TUB were the most stable reference genes recommended by geNorm, COPS3, ABCC2, R3HDM2 by NormFinder, CAC, UBC2 and DNAJ by BestKeeper, COPS3, R3HDM2 and ABCC2 by Delta CT (Table 3).
Because the growth, development, and metabolite accumulation of all the living organisms are certainly influenced by multiple factors, therefore, we also studied the optimal reference genes under the spatial–temporal interaction conditions in both G. uralensis and G. inflata. In G. uralensis, the top three most stable candidate reference genes were CAC, R3HDM2, ABCC2 identified by geNorm, DNAJ, CAC, R3HDM2 by NormFinder, COPS3, TIF1, UBC2 by BestKeeper, and CAC, R3HDM2, DNAJ by Delta CT (Table 2). In G. inflata, DREB, ABCC2and CS were the most stable reference genes recommended by geNorm, COPS3, DREB and CAC by NormFinder and Delta CT, ABCC2, COPS3 and DREB by BestKeeper (Table 3).
Selection of optimal reference genes under different experimental conditions
Based on RefFinder, a web-based software, comprehensive ranking of reference genes integrating all four software was obtained (Tables 2 and 3). At different experimental stages, CAC, CYP, ABCC2 were identified as the top three most stable reference genes in the G. uralensis (Fig. 3A, Table 2), while RAN, COPS3, CS were identified in G. inflata (Fig. 3B, Table 3). For the different tissues, R3HDM2, CAC, TUB were identified as the most stable reference genes in the G. uralensis (Fig. 3A, Table 2), while COPS3, R3HDM2, DREB were identified in the G. inflata (Fig. 3B, Table 3). Under the spatial–temporal interaction conditions in G. uralensis, CAC, R3HDM2, DNAJ were identified as the most stable reference genes in the G. uralensis (Fig. 3A, Table 2), while COPS3, DREB, ABCC2 were identified in the G. inflata (Fig. 3B, Table 3).
The optimal number of reference genes under each experimental conditions required for reliable normalization in two species were predicted by geNorm software with the V values (cutoff = 0.15). When pairwise variations Vn/n + 1 < 0.15, it means that an addition reference gene (n + 1) is not necessary. For all the experimental conditions in G. uralensis the first V-value less than 0.15 occurred at V2/3, suggesting that two reference genes were adequate to correctly normalize gene expression. But for spatial–temporal interaction conditions in the G. inflata, more than two reference genes was necessary suggested by V-value for accurate normalization, the first V-value less than 0.15 occurred at V4/5 (Fig. 4).
Pairwise variation analysis of the candidate reference genes. Pairwise variation analysis in G. uralensis and G. inflata, respectively, by geNorm procedure. The pairwise variation (Vn/Vn+1) was calculated with the normalization factors NFn and NFn+1 to determine the optimal number of reference genes required for RT-qPCR data normalization under different conditions.
Comparison of the suitable reference genes under different experimental conditions and between the two congenic Glycyrrhiza species
The summary of the top three most suitable reference genes under all experimental condition showed that seven genes appeared in the top three list in G. uralensis, among which CAC and R3HDM2 showed the highest recommended frequency (33.33% and 22.22%, respectively) (Fig. 3A). In G. inflata, seven genes were also appeared in the top three lists, and COPS3 and DREB presented the highest recommended frequency, with the frequency of 33.33% and 22.22%, respectively (Fig. 3B).
Comparison of the suitable reference genes under different experimental conditions showed only one of the top three candidates was shared between the individual factors and their interactions, specifically, CAC in G. uralensis and COPS3 in G. inflata. Therefore, CAC was the most stable reference gene in G. uralensis under all the experimental conditions tested, while COPS3 was the most stable in G. inflata.
For the comparison of the suitable reference genes between the two congenic Glycyrrhiza species, we found R3HDM2 was the only suitable reference gene shared between G. uralensis and G. inflata in different tissues, and no consistent reference gene was found under different developmental stage and tissue and developmental stage interactions between the two congenic Glycyrrhiza species. So the optimal reference genes for different species are variable, even for the two proximal species in the same genus.
Validation of recommended reference genes
A root and rhizome-specific gene, β-AS, a key gene in glychrizin biosynthesis was used to validate the selected reference genes. To validate the reference geneto study expressin pattern in different tissues, expression of β-AS was normalized to both the most and the least stablecandidate reference genes, both in G.uralensis and G. inflata. When the recommended reference genes were used, the expression levels of β-AS in the roots and the rhizomes were similar and both high, while its expression was significantly reduced in the leaf. However, when normalized to the least suited reference genes, the expression pattern of β-AS changed, or the ratio of expression levels between roots/rhizomes and leaves were significantly enlarged or narrowed (Fig. 5). Unstable reference genes really confuse the results.
Validation of the recommended reference genes. Expression profiles of β-AS gene among different tissues were investigated using seven different normalization factors. The most suited, the top two most suited, the top three most suited, the least suited, the top two least suited, the top three least suited, and all the candidate reference genes. Bars represent the means ± standard error of biological replications. Different letters indicate significant differences (P < 0.05).
Survey of the reference genes used within Leguminosae plants
Reference gene selection has been reported in 12 Leguminosae species (Arachis hypogaea L., Cassia obtusifolia L., Cicer arietinum L., Cyamopsis tetragonoloba L.Taub, Eremosparton songoricum (Litv.) Vass., Glycine max (L.) Merr., Hedysarum coronarium L., Lens culinaris Medic., Lupinus angustifolius L., Medicago sativa L., Phaseolus vulgaris Linn., Vigna angularis (Willd.) Ohwi et Ohashi) under different experimental conditions, including different tissues or different experimental stages. In this study, we added G. uralensis and G. inflata to this list.
A total of ten species (including G. uralensis and G. inflata) had been studied under different developmental stages, among the 22 reference genes recommended, eukaryotic elongation factor (EF, EF1α, and ELF1B) and tubulin (TUA1, TUA2, TUA5, and TUB) were the most choices, and the frequency of recommendation in the 10 species were 15.15% and 12.12%, respectively (Fig. 6). For the different tissues, a total of twelve species had been studied. Among the 20 reference genes recommended, ACT, EF, and UBQ performed particularly well, and they presented recommended frequency of 16.66%, 14.58% and 12.50%, respectively (Fig. 6).
Frequency of reference genes recommended for RT-qPCR analysis under different conditions in Leguminosae plants. Here we surveyed the frequency of each reference gene recommended for different developmental stages and different tissues among the twelve Leguminosae species. Reference genes (top three) recommended for each species under different developmental stages and tissues are detailed in Table S5.
Discussion
Licorice are herbaceous perennial plants with great medical and ecological values. Licorice root extracts have been proved to have anti-carcinogenic2,3, anti-inflammatory, anti-fungal, anti-piroplasmic and cytotoxic activities4. More recently, glycyrrhizin, the most important bioactive triterpenoid saponin in licorice roots, is under the consideration for treating COVID-19 infection caused respiratory syndrome5. Accumulation of its many bioactive compounds is spatio-temporal dependent. To study the functional genes involved in development and biosynthesis of these bioactive compounds, spatio-temporal expression pattern of these genes provides an important piece of information. Here we report a set of reference genes for RT-qPCR analysis to study spatio-temporal expression pattern of genes in two congenic licorice species, G. uralensis and G. inflata.
Candidates screening and optimal reference genes selection
In this study, CAC and COPS3 was one of the top candidates for all three experimental conditions tested in G. uralensis and G. inflata, respectively. COPS3 is a new candidate of reference gene screened from RNA-seq, and it was proved to be the most stable gene under different developmental stages in G. inflata. This result supports the idea that it is feasible to screen reference genes through RNA-seq dataset35. The current and emerging RNA-seq data may provide a bigger and even more reliable pool to select candidate reference genes other than the traditional housekeeping genes. Such discoveries are of great significance and should enable greater accuracy of normalization, particularly across diverse plant organs and in other experimental conditions where traditional housekeeping genes display variability in expression.
The suitable reference genes for spatio-temporal gene expression is different from that for individual conditions
Our results showed that different reference genes were selected to study spatial, temporal expression as well as spatial–temporal expression patterns in both species. The most stable reference genes varied in different experiments. In G. uralensis, the top three most stable reference genes across different tissues were R3HDM2, CAC and TUB, while CAC, CYP and ABCC2 were most suited for different developmental stages. Similarly, in G. inflata, COPS3, R3HDM2 and DREB were selected for tissues, while RAN, COPS3 and CS were recommended for developmental stages (Tables 2 and 3). In addition, the optimal reference genes under the condition of two-factor interaction (developmental stages × tissues) are also different from those under single factor conditions. For the tissue-developmental stage interactions, CAC, R3HDM2 and DNAJ were the most suited reference genes in G. uralensis, while COPS3, DREB and ABCC2 in G. inflata. Only one of the top three candidates was shared between the individual factors and their interactions, specifically, CAC in G. uralensis and COPS3 in G. inflata (Fig. 3). Because the expression of genes is constantly under the influences of multiple factors/dimensions, so it is essential for gene function analysis to investigate gene expression under the interacting factors. Our results in this study illustrated that the optimal reference gene for spatio-temporal gene expression is different from that for individual conditions, so every gene expression analysis should begin with validation of reference genes in a given sample set under specific experiment conditions, either under single factor or under multiple factors interacting conditions.
The validation of selected reference genes was done by normalizing the expression of β-AS, a key gene involved in glycyrrhizin biosynthesis, and its expression has been proven to be mainly in roots and rhizomes8,9. Generally, the expression pattern of β-AS should not be affected by reference gene selection, because the M values of the 14 candidate genes selected in our study were all below 1.5 by geNorm. However, our results showed that the expression pattern of β-AS was quite different when using the unstable reference genes for homogenization compared with the stable reference genes (Fig. 5). Therefore, our results showed that unstable reference genes would confuse the expression pattern while the stable reference genes gave reliable results, and the optimal reference genes screened in this study are reliable.
The optimal reference genes within Leguminosae species are drastically different
In this study, we also summarized the validated reference genes for different development stages in Leguminosae (including G. uralensis and G. inflata tested in this study). From the results of our survey, we found EF1, TUB and UBQ are commonly used housekeeping genes, which have been identified as the most suitable ones at different developmental stages in several species. EF had been selected as the most suitable reference gene at different developmental stages in E. songoricum36, G. max37, and C. arietinum38. TUB was the optimal reference gene at different experimental stages in E. songoricum36, G. max37, and H. coronarium39. While UBQ was validated as the most stable reference gene at different developmental stages in C. tetragonoloba40, H. coronarium39, L. angustifolius41. For the different tissues in Leguminosae, the most stable reference gene candidates were ACT, EF and UBQ. Among them, ACT was recommended as a stable reference gene in different tissues of C. tetragonoloba40, E. songoricum36, G. max37,42, P. vulgaris43. EF was recommended in C. obtusifolia44, G. max37, M. sativa45, and UBQ was recommended in G. inflata (Table 3), E. songoricum 36, G. max37, and L. angustifolius41. So, the optimal reference genes for different species in the same family are variable, even for the two proximal species in the same genus (G. uralensis and G. inflata). We found CAC was the most stable reference gene when all factors taken account in G. uralensis, and COPS3 was the optimal reference gene with the highest recommended frequency in G. inflata (Fig. 3). However, EF1 and TIF1 in G. uralensis, and ACT and TIF1 G. inflata were the most unstable reference genes respectively, and it has been proved that it will cause false results using the unstable reference gene for expression normalization (Fig. 5). Among them, EF1 and ACT are commonly used housekeeping gene and have been identified as the most suitable reference genes in several studies40,43,46. So our results indicated that it is always necessary to validate reference genes for reliable gene expression analysis. The summary and analysis of the reported legume reference genes will serves as a guide for the subsequent selection of reference genes in Leguminosae plants.
Summary and perspectives
In this study, we evaluated the expression of 14 candidate reference genes across different tissues (root, rhizome, leaf) at various developmental stages (returning green, April; rapid growth and flowering, May; seed setting, July; and senescence stage, October) in the two congeneric medicinal plants, G. uralensis and G. inflate, respectively. Based on previous research and our preliminary data, we hypothesized that internal references for spatio-temporal gene expression are different from the reference genes suited for individual factors. In G. uralensis, the top three most stable reference genes across different tissues were R3HDM2, CAC and TUB, while CAC, CYP and ABCC2 were most suited for different developmental stages. CAC is the only candidate recommended for both biotic factors, which is reflected in the stability ranking for the spatio (tissue)-temporal (developmental stage) interactions (CAC, R3HDM2 and DNAJ). Similarly, in G. inflata, COPS3, R3HDM2 and DREB were selected for tissues, while RAN, COPS3 and CS were recommended for developmental stages. For the tissue-developmental stage interactions, COPS3, DREB and ABCC2 were the most suited reference genes. In both species, only one of the top three candidates was shared between the individual factors and their interactions, specifically, CAC in G. uralensis and COPS3 in G. inflata, which supports our overarching hypothesis.
In addition, we also documented the reference genes that have been used in RT-qPCR analyses among 12 different Leguminosae plants under the same biotic conditions with current study, i.e., tissue and/or developmental stage. Among the 115 genes have been tested, even the routinely used reference genes showed variable expressions under different experimental conditions. Therefore, to avoid the misinterpretation of RT-qPCR results, a thorough evaluation of reference genes is strongly recommended. More importantly, given that biosynthesis of bioactive compounds is typically spatio-temporal dependent, the selection of suitable reference genes should follow suit. Based on previous studies and our current results, we concluded (1) transcriptome is a rich reservoir for selecting stably expressed candidate reference genes, (2) customized design, especially the interaction among the experimental conditions, is warranted for searching suitable reference genes in any given species, and (3) without validation study, gene(s), including housekeeping genes, could lead to ambiguous results, especially in non-model species. Finally, spatio-temporal selection of reference genes not only lays the foundation for functional genomics research in Glycyrrhiza, but also facilitates these traditional medicinal herbs to reach/maximize their pharmaceutical potential.
Data availability
Data will be available upon request. RNA-seq datasets used in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found below: https://www.ncbi.nlm.nih.gov/, PRJNA574093.
Abbreviations
- ABCC2:
-
ATP binding-box transporter 2
- ACT:
-
Actin1 gene
- CAC:
-
Clathrin complex AP1
- COPS3:
-
COP9 signal complex subunit 3
- CS:
-
Citrate synthase
- CYP:
-
Cyclophilin
- CYP88D6:
-
β-Amyrin 11-oxidase
- DNAJ:
-
Heat-shock protein 40
- DREB:
-
Dehydration responsive element binding gene
- E:
-
The efficiency of the PCR amplification
- EF1:
-
Translation elongation factor1
- MIQE:
-
Minimum information for publication of quantitative real-time PCR experiments
- RT-qPCR:
-
Reverse transcription quantitative real-time PCR
- R3HDM2:
-
R3H domain protein 2
- RAN:
-
Ras related protein
- TIF1:
-
Translation initiation factor
- TUB:
-
β-Tubulin
- UBC2:
-
Ubiquitin-conjugating enzyme E2
- β-AS:
-
β-Amyrin synthase
References
Gibson, M. R. Glycyrrhiza in old and new perspectives. Lloydia 41, 348–354 (1978).
Hao, W. et al. Licochalcone A-induced human gastric cancer BGC-823 cells apoptosis by regulating ROS-mediated MAPKs and PI3K/AKT signaling pathways. Sci. Rep. 5, 10336. https://doi.org/10.1038/srep10336 (2015).
Tang, Z. H. et al. Induction of C/EBP homologous protein-mediated apoptosis and autophagy by licochalcone A in non-small cell lung cancer cells. Sci. Rep. 6, 26241. https://doi.org/10.1038/srep26241 (2016).
Batiha, G., Beshbishy, A. & Abdel Daim, M. Traditional uses, bioactive chemical constituents, and pharmacological and toxicological activities of the miracle medicinal herb; Glycyrrhiza glabra L. (Fabaceae Family). Biomolecules 10, 352. https://doi.org/10.3390/biom10030352 (2020).
Bailly, C. & Vergoten, G. Glycyrrhizin: An alternative drug for the treatment of COVID-19 infection and the associated respiratory syndrome?. Pharmacol. Therap. 214, 107618. https://doi.org/10.1016/j.pharmthera.2020.107618 (2020).
Hayashi, H. & Sudo, H. Economic importance of licorice. Plant Biotechnol. 26, 101–104. https://doi.org/10.5511/plantbiotechnology.26.101 (2009).
Souri, M. Changes in glycyrrhizin content of iranian licorice (Glycyrrhiza glabra L.) affected by different root diameter and ecological conditions. Agric. Commun. 2, 27–33 (2014).
Seki, H. et al. Licorice β-amyrin 11-oxidase, a cytochrome P450 with a key role in the biosynthesis of the triterpene sweetener glycyrrhizin. Proc Natl Acad Sci USA 105, 14204–14209. https://doi.org/10.1073/pnas.0803876105 (2008).
Seki, H. et al. Triterpene functional genomics in licorice for identification of CYP72A154 involved in the biosynthesis of glycyrrhizin. Plant Cell 23, 4112–4123. https://doi.org/10.1105/tpc.110.082685 (2011).
Tamura, K. et al. The basic helix-loop-helix transcription factor GubHLH3 positively regulates soyasaponin biosynthetic genes in Glycyrrhiza uralensis. Plant Cell Physiol. 59, 783–793. https://doi.org/10.1093/pcp/pcy046 (2018).
Simpson, G. G. & Dean, C. Arabidopsis, the Rosetta stone of flowering time?. Science (New York, NY) 296, 285–289. https://doi.org/10.1126/science.296.5566.285 (2002).
Smith, Z. R. & Long, J. A. Control of Arabidopsis apical-basal embryo polarity by antagonistic transcription factors. Nature 464, 423–426. https://doi.org/10.1038/nature08843 (2010).
Christensen, L. P. Ginsenosides: Chemistry, biosynthesis, analysis, and potential health effects. Adv. Food Nutr. Res. 55, 1–99 (2008).
Han, J. Y., Hwang, H. S., Choi, S. W., Kim, H. J. & Choi, Y. E. Cytochrome P450 CYP716A53v2 catalyzes the formation of protopanaxatriol from protopanaxadiol during ginsenoside biosynthesis in Panax ginseng. Plant Cell Physiol. 53, 1535–1545. https://doi.org/10.1093/pcp/pcs106 (2012).
Carlson, M. R. et al. Gene connectivity, function, and sequence conservation: predictions from modular yeast co-expression networks. BMC Genomics 7, 40. https://doi.org/10.1186/1471-2164-7-40 (2006).
Stuart, J. M., Segal, E., Koller, D. & Kim, S. K. A gene-coexpression network for global discovery of conserved genetic modules. Science (New York, NY) 302, 249–255. https://doi.org/10.1126/science.1087447 (2003).
Phillips, M. A., D’Auria, J. C., Luck, K. & Gershenzon, J. Evaluation of candidate reference genes for real-time quantitative PCR of plant samples using purified cDNA as template. Plant Mol. Biol. Rep. 27, 407–416. https://doi.org/10.1007/s11105-008-0072-1 (2009).
Udvardi, M. K., Czechowski, T. & Scheible, W.-R. Eleven golden rules of quantitative RT-PCR. Plant Cell 20, 1736–1737. https://doi.org/10.1105/tpc.108.061143 (2008).
Hong, S. Y., Seo, P. J., Yang, M., Xiang, F. & Park, C. Exploring valid reference genes for gene expression studies in Brachypodium distachyonby real-time PCR. BMC Plant Biol. 8, 112–112. https://doi.org/10.1186/1471-2229-8-112 (2008).
Dheda, K. Validation of housekeeping genes for normalizing RNA expression in real-time PCR. Biotechniques 37, 118–119. https://doi.org/10.1016/j.biosystems.2004.03.004 (2004).
Nicot, N., Hausman, J. F., Hoffmann, L. & Evers, D. Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J. Exp. Bot. 56, 2907–2914. https://doi.org/10.1093/jxb/eri285 (2005).
Meller, M., Vadachkoria, S., Luthy, D. A. & Williams, M. A. Evaluation of housekeeping genes in placental comparative expression studies. Placenta 26, 601–607. https://doi.org/10.1016/j.placenta.2004.09.009 (2005).
Wang, M., Wang, Q. & Zhang, B. Evaluation and selection of reliable reference genes for gene expression under abiotic stress in cotton (Gossypium hirsutum L.). Gene 530, 44–50. https://doi.org/10.1016/j.gene.2013.07.084 (2013).
Bustin, S. A. Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): trends and problems. Clin. Chem. 55, 611–622. https://doi.org/10.1373/clinchem.2008.112797 (2009).
Hayashi, H., Hiraoka, N., Ikeshiro, Y., Yamamoto, H. & Yoshikawa, T. Seasonal variation of glycyrrhizin and isoliquiritigenin glycosides in the root of Glycyrrhiza glabra L. Biol. Pharm. Bull. 21, 987–989. https://doi.org/10.1248/bpb.21.987 (1998).
Ramilowski, J. A. et al. Glycyrrhiza uralensis transcriptome landscape and study of phytochemicals. Plant Cell Physiol. 54, 697–710. https://doi.org/10.1093/pcp/pct057 (2013).
Hayashi, H. et al. Cloning and characterization of a cDNA encoding beta-amyrin synthase involved in glycyrrhizin and soyasaponin biosyntheses in licorice. Biol. Pharm. Bull. 24, 912–916. https://doi.org/10.1248/bpb.24.912 (2001).
Maroufi, A. Selection of reference genes for real-time quantitative PCR analysis of gene expression in Glycyrrhiza glabra under drought stress. Biol. Plantarum 60, 1–10. https://doi.org/10.1007/s10535-016-0601-y (2016).
Zeng, S. et al. Identification and validation of reference genes for quantitative teal-time PCR normalization and Its applications in Lycium. PLoS ONE 9, e97039. https://doi.org/10.1371/journal.pone.0097039 (2014).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, 1–12. https://doi.org/10.1186/gb-2002-3-7-research0034 (2002).
Andersen, C. L., Jensen, J. L. & Ørntoft, T. F. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 64, 5245–5250. https://doi.org/10.1158/0008-5472.can-04-0496 (2004).
Pfaffl, M. W., Tichopad, A., Prgomet, C. & Neuvians, T. P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—excel-based tool using pair-wise correlations. Biotechnol. Lett. 26, 509–515. https://doi.org/10.1023/b:bile.0000019559.84305.47 (2004).
Arocho, A., Chen, B., Ladanyi, M. & Pan, Q. Validation of the 2-DeltaDeltaCt calculation as an alternate method of data analysis for quantitative PCR of BCR-ABL P210 transcripts. Diagn. Mol. Pathol. 15, 56–61. https://doi.org/10.1097/00019606-200603000-00009 (2006).
Xie, F., Xiao, P., Chen, D., Xu, L. & Zhang, B. miRDeepFinder: a miRNA analysis tool for deep sequencing of plant small RNAs. Plant Mol. Biol. 80, 75–84. https://doi.org/10.1007/s11103-012-9885-2 (2012).
Czechowski, T., Stitt, M., Altmann, T., Udvardi, M. K. & Scheible, W.-R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 139, 5–17. https://doi.org/10.1104/pp.105.063743 (2005).
Li, X. S., Yang, H. L., Zhang, D. Y., Zhang, Y. M. & Wood, A. J. Reference gene selection in the desert plant Eremosparton songoricum. Int. J. Mol. Sci. 13, 6944–6963. https://doi.org/10.3390/ijms13066944 (2012).
Miranda Vde, J. et al. Validation of reference genes aiming accurate normalization of qPCR data in soybean upon nematode parasitism and insect attack. BMC Res. Notes 6, 196. https://doi.org/10.1186/1756-0500-6-196 (2013).
Reddy, D. S. et al. Identification and validation of reference genes and their impact on normalized gene expression studies across cultivated and wild Cicer species. PLoS ONE https://doi.org/10.1371/journal.pone.0148451 (2016).
Cordoba, E. M., Die, J. V., Gonzalez-Verdejo, C. I., Nadal, S. & Roman, B. Selection of reference genes in Hedysarum coronarium under various stresses and stages of development. Anal. Biochem. 409, 236–243. https://doi.org/10.1016/j.ab.2010.10.031 (2011).
Jaiswal, P. S., Kaur, N. & Randhawa, G. S. Identification of reference genes for qRT-PCR gene expression studies during seed development and under abiotic stresses in Cyamopsis tetragonoloba. Crop. Sci. 59, 252–265. https://doi.org/10.2135/cropsci2018.05.0313 (2019).
Taylor, C. M., Jost, R., Erskine, W. & Nelson, M. N. Identifying stable reference genes for qRT-PCR normalisation in gene expression studies of narrow-leafed Lupin (Lupinus angustifolius L.). Plos One 11, e0148300. https://doi.org/10.1371/journal.pone.0148300 (2016).
Hu, R., Fan, C., Li, H., Zhang, Q. & Fu, Y.-F. Evaluation of putative reference genes for gene expression normalization in soybean by quantitative real-time RT-PCR. BMC Mol. Biol. 10, 93. https://doi.org/10.1186/1471-2199-10-93 (2009).
Pereira, W. J., Bassinello, P. Z., Brondani, C. & Vianello, R. P. An improved method for RNA extraction from common bean seeds and validation of reference genes for qPCR. Crop Breed. Appl. Biotechnol. 17, 150–158. https://doi.org/10.1590/1984-70332017v17n2a22 (2017).
Liu, Z. et al. Selection and evaluation of reference genes for expression analysis of Cassi. Biosci. Biotechnol. Biochem. 79, 1818–1826. https://doi.org/10.1080/09168451.2015.1052771 (2015).
Wang, X. et al. Selection of reliable reference genes for quantitative real-time RT-PCR in alfalfa. Genes Genet. Syst. 90, 175–180. https://doi.org/10.1266/ggs.90.175 (2015).
Chi, X. et al. Validation of reference genes for gene expression studies in peanut by quantitative real-time RT-PCR. Mol. Genet. Genom. 287, 167–176. https://doi.org/10.1007/s00438-011-0665-5 (2012).
Acknowledgements
This work was supported by National Key R&D Project from the Ministry of Science and Technology of China (2019YFC1711102), National Natural Science Foundation of China (31270319), Department of Science and Technology of Guangzhou (202002030442) and Guangdong Provincial Special Fund For Modern Agriculture Industry Technology Innovation Teams, China (2019KJ148).
Author information
Authors and Affiliations
Contributions
This research was designed by Y.L., X.Z., and Y.W., Y.L. and X.L. carried out the experiments. Y.L. and X.Z. performed data analysis. M.L. and Y.A. collected samples from Ningxia. Y.L. drafted and Y.L., X.Z., and L.Y. revised the manuscript. All authors read and approved its final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, Y., Liang, X., Zhou, X. et al. Spatio-temporal selection of reference genes in the two congeneric species of Glycyrrhiza. Sci Rep 11, 1122 (2021). https://doi.org/10.1038/s41598-020-79298-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-79298-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.