Abstract
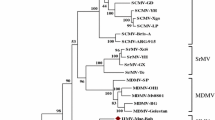
The host range of previously reported bymoviruses is restricted to plants belonging to the family Poaceae. Soybean leaf rugose mosaic virus (SbLRMV) from non-Poaceae plants is related to bymoviruses based on a partial genome sequence. However, unlike bymoviruses, this virus infects plants of at least four dicotyledonous families, including Fabaceae, and causes disease in soybean. Complete nucleotide sequences of two variants of SbLRMV were determined, and its taxonomic position was clarified. RNA1 is 7109 nucleotides (nt) long with one large open reading frame (ORF), possibly encoding a polyprotein of 257 kDa. This polyprotein is likely processed into eight mature proteins. The entire RNA1 ORF shares 52%–55% nucleotide sequence identity and 27%-43% amino acid sequence identity, and the coat protein shares 49%–54% nucleotide sequence identity and 30%–34% amino acid sequence identity to other bymoviruses. The similarity to other viruses in the family Potyviridae is generally lower. RNA2 is 3413 or 3415 nt long and putatively encodes a polyprotein of 108 kDa. This protein is probably cleaved into two mature proteins. The sequences of these two RNAs are very similar to those of bymoviruses. Phylogenetic analysis of members of the family Potyviridae showed that RNA1 and RNA2 of SbLRMV formed a basal clade with known bymoviruses. Inoculation tests using leaf samples suggested that SbLRMV RNA1 can systemically infect and cause disease in soybean without the presence of RNA2. In conclusion, SbLRMV is an atypical member of the genus Bymovirus that infects soybean (Fabaceae) and other dicots rather than gramineous hosts.



Similar content being viewed by others
References
Kuroda T, Nabata K, Hori T et al (2010) Soybean leaf rugose mosaic virus, a new soilborne virus in the family Potyviridae, isolated from soybean in Japan. J Gen Plant Pathol 76:382–388
Adams MJ, Zerbini FM, French R et al (2012) Bymovirus. In: King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ (eds) Virus taxonomy. Ninth Report of the International Committee on Taxonomy of Viruses. Elsevier/Academic Press, San Diego, pp 1084–1086
Zheng T, Chen J, Chen JP, Adams MJ (2002) The complete sequence of Oat mosaic virus and evidence for deletion and duplication in RNA2. Arch Virol 147:635–642
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Namba S, Kashiwazaki S, Lu X et al (1998) Complete nucleotide sequence of wheat yellow mosaic bymovirus genomic RNAs. Arch Virol 143:631–643
Wagh SG, Kobayashi K, Yaeno T et al (2016) Rice necrosis mosaic virus, a fungal transmitted Bymovirus: complete nucleotide sequence of the genomic RNAs and subgrouping of bymoviruses. J Gen Plant Pathol 82:38–42
Kadaré G, Haenni A-L (1997) Virus-encoded RNA helicases. J Virol 71:2583–2590
Chung BYW, Miller WA, Atkins JF, Firth AE (2008) An overlapping essential gene in the Potyviridae. Proc Natl Acad Sci USA 105:5897–5902
Kashiwazaki S (1996) The complete nucleotide sequence and genome organization of barley mild mosaic virus (Nal strain). Arch Virol 141:2077–2089
Kashiwazaki S, Minobe Y, Hibino H (1991) Nucleotide sequence of barley yellow mosaic virus RNA 2. J Gen Virol 72:995–999
Adams MJ, Antoniw JF, Fauquet CM (2005) Molecular criteria for genus and species discrimination within the family Potyviridae. Arch Virol 150:459–479
You Y, Shirako Y (2010) Bymovirus reverse genetics: requirements for RNA2-encoded proteins in systemic infection. Mol Plant Pathol 11:383–394
Jacobi V, Peerenboom E, Schenk PM et al (1995) Cloning and sequence analysis of RNA-2 of a mechanically transmitted UK isolate of barley mild mosaic bymovirus (BaMMV). Virus Res 37:99–111
Peerenboom E, Jacobi V, Antoniw JF et al (1996) The complete nucleotide sequence of RNA-2 of a fungally-transmitted UK isolate of barley mild mosaic bymovirus and identification of amino acid combinations possibly involved in fungus transmission. Virus Res 40:149–159
Kanyuka K, Ward E, Adams MJ (2003) Polymyxa graminis and the cereal viruses it transmits: a research challenge. Mol Plant Pathol 4:393–406
Legrève A, Delfosse P, Maraite H (2002) Phylogenetic analysis of Polymyxa species based on nuclear 5.8S and internal transcribed spacers ribosomal DNA sequences. Mycol Res 106:138–147
Delfosse P, Reddy AS, Thirumala Devi K et al (2002) Dynamics of Polymyxa graminis and Indian peanut clump virus (IPCV) infection on various monocotyledonous crops and groundnut during the rainy season. Plant Pahtol 51:546–560
Nakamura S, Tomita S, Honkura R, Seo N (2016) Characterization of a filamentous virus from broad bean showing chlorotic spot symptoms (Abstract in Japanese). Jpn J Phytopathol 82:253
Tomitaka Y, Usugi T, Kunitomo E et al (2014) A necrosis and mosaic disease by a novel bymo-like virus occurred on broad bean (Abstract in Japanese). Jpn J Phytopathol 80:16
Funding
No funding was received.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Takehiro Ohki, Tomohisa Kuroda, Mitsuru Sayama, and Tetsuo Maoka declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: Ralf Georg Dietzgen.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ohki, T., Kuroda, T., Sayama, M. et al. Complete nucleotide sequence of soybean leaf rugose mosaic virus, an atypical member of the genus Bymovirus. Arch Virol 166, 1885–1892 (2021). https://doi.org/10.1007/s00705-021-05069-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05069-z