Abstract
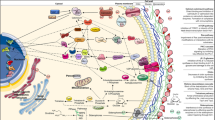
Treating fungal infections is challenging and frequently requires long-term courses of antifungal drugs. Considering the limited number of existing antifungal drugs, it is crucial to evaluate the possibility of repositioning drugs with antifungal properties and to revisit older antifungals for applications in combined therapy, which could widen the range of therapeutic possibilities. Undecanoic acid is a saturated medium-chain fatty acid with known antifungal effects; however, its antifungal properties have not been extensively explored. Recent advances indicate that the toxic effect of undecanoic acid involves modulation of fungal metabolism through its effects on the expression of fungal genes that are critical for virulence. Additionally, undecanoic acid is suitable for chemical modification and might be useful in synergic therapies. This review highlights the use of undecanoic acid in antifungal treatments, reinforcing its known activity against dermatophytes. Specifically, in Trichophyton rubrum, against which the activity of undecanoic acid has been most widely studied, undecanoic acid elicits profound effects on pivotal processes in the cell wall, membrane assembly, lipid metabolism, pathogenesis, and even mRNA processing. Considering the known antifungal activities and associated mechanisms of undecanoic acid, its potential use in combination therapy, and the ability to modify the parent compound structure, undecanoic acid shows promise as a novel therapeutic against fungal infections.



Similar content being viewed by others
References
Martinez-Rossi NM, Bitencourt TA, Peres NTA, Lang EAS, Gomes EV, Quaresemin NR, et al. Dermatophyte resistance to antifungal drugs: mechanisms and prospectus. Front Microbiol. 2018;9:1108.
Cordoba S, Vivot W, Szusz W, Albo G. Antifungal activity of essential oils against Candida species isolated from clinical samples. Mycopathologia. 2019;184(5):615–23.
Lopes AI, Tavaria FK, Pintado ME. Conventional and natural compounds for the treatment of dermatophytosis. Med Mycol. 2020;58(6):707–20.
Souza ACO, Amaral AC. Antifungal therapy for systemic mycosis and the nanobiotechnology era: improving efficacy, biodistribution and toxicity. Front Microbiol. 2017;8:336.
Drake DR, Brogden KA, Dawson DV, Wertz PW. Thematic review series: skin lipids—antimicrobial lipids at the skin surface. J Lipid Res. 2008;49(1):4–11.
Feingold KR. The outer frontier: the importance of lipid metabolism in the skin. J Lipid Res. 2009;50:S417–22.
Clark JF. On the toxic effect of deleterious agents on the germination and development of certain filamentous fungi. Bot Gaz. 1899;28(5):289–327.
Keeney EL, Ajello L, Broyles EN, Lankford E. Propionate and undecylenate ointments in the treatment of tinea pedis and an in vitro comparison of their fungistatic and antibacterial effects with other ointments. Bull Johns Hopkins Hosp. 1944;74:417–39.
Shapiro AL, Rothman S. Undecylenic acid in the treatment of dermatomycosis. Arch Dermatol. 1945;119(4):345–50.
Peck SM, Russ WR. Propionate-caprylate mixtures in the treatment of dermatomycoses, with a review of fatty acid therapy in general. Arch Derm Syphilol. 1947;56(5):601–13.
Kiesel A. Recherches sur L’action de divers acides et sels acides sur le développement de L`Aspergillus niger. Ann Inst Pasteur. 1913;27:391–420.
Hoffman C, Schweitzer TR, Dalby G. Fungistatic properties of the fatty acids and possible biochemical significance. J Food Sci. 1939;4(6):539–45.
Peck SM, Rosenfeld H. The effects of hydrogen ion concentration, fatty acids and vitamin C on the growth of fungi. J Invest Dermatol. 1938;1(4):237–65.
Garg AP, Muller J. Fungitoxicity of fatty acids against dermatophytes. Mycoses. 1993;36(1–2):51–63.
Vicher EE, Lyon I, White EL. Studies on the respiration of Trichophyton rubrum. Mycopathol Mycol Appl. 1959;11:185–95.
Vicher EE, Kostiw LL, Lyon I, Bolewicz BM. The effect of sodium propionate and sodium caprylate on the fatty acid content of Trichophyton rubrum. Mycopathol Mycol Appl. 1968;35(3):208–14.
Samson FE, Katz AM, Harris DL. Effects of acetate and other short-chain fatty acids on yeast metabolism. Arch Biochem Biophys. 1955;54(2):406–23.
Das SK, Adhya S, Banerjee AB. Effect of undecanoic acid on germination of microconidia of wild and undecanoic acid resistant mutant of Trichophyton rubrum. Mycopathologia. 1977;61(2):121–3.
Das SK, Banerjee AB. Effect of undecanoic acid on phospholipid-metabolism in Trichophyton rubrum. Sabouraudia-J Med Vet Mycol. 1982;20(4):267–72.
Das SK, Banerjee AB. Effect of undecanoic acid on the production of exocellular lipolytic and keratinolytic enzymes by undecanoic acid-sensitive and -resistant strains of Trichophyton rubrum. Sabouraudia. 1982;20(3):179–84.
Das SK, Banerjee AB. Effect of undecanoic acid on lipid composition of Trichophyton rubrum. Mycopathologia. 1983;83(1):35–9.
Peres NTA, Cursino-Santos JR, Rossi A, Martinez-Rossi NM. In vitro susceptibility to antimycotic drug undecanoic acid, a medium-chain fatty acid, is nutrient-dependent in the dermatophyte Trichophyton rubrum. World J Microbiol Biotechnol. 2011;27(7):1719–23.
Martins MP, Silva LG, Rossi A, Sanches PR, Souza LDR. Martinez-Rossi NM. Global analysis of cell wall genes revealed putative virulence factors in the dermatophyte Trichophyton rubrum. Front Microbiol. 2019;10:2168.
Mendes NS, Bitencourt TA, Sanches PR, Silva-Rocha R, Martinez-Rossi NM, Rossi A. Transcriptome-wide survey of gene expression changes and alternative splicing in Trichophyton rubrum in response to undecanoic acid. Sci Rep. 2018;8(1):2520.
Gomes EV, Bortolossi JC, Sanches PR, Mendes NS, Martinez-Rossi NM, Rossi A. STE20/PAKA protein kinase gene releases an autoinhibitory domain through pre-mRNA alternative splicing in the dermatophyte Trichophyton rubrum. Int J Mol Sci. 2018;19(11):3654.
Neves-da-Rocha J, Bitencourt TA, de Oliveira VM, Sanches PR, Rossi A, Martinez-Rossi NM. Alternative splicing in heat shock protein transcripts as a mechanism of cell adaptation in Trichophyton rubrum. Cells. 2019;8(10):1206.
Avrahami D, Shai Y. Bestowing antifungal and antibacterial activities by lipophilic acid conjugation to D, L-amino acid-containing antimicrobial peptides: a plausible mode of action. Biochemistry. 2003;42(50):14946–56.
Muthamil S, Balasubramaniam B, Balamurugan K, Pandian SK. Synergistic effect of quinic acid derived from Syzygium cumini and undecanoic acid against Candida spp. biofilm and virulence. Front Microbiol. 2018;9:2835.
Lee JH, Kim YG, Khadke SK, Lee J. Antibiofilm and antifungal activities of medium-chain fatty acids against Candida albicans via mimicking of the quorum-sensing molecule farnesol. Microb Biotechnol. 2020.
Hastings J, Owen G, Dekker A, Ennis M, Kale N, Muthukrishnan V, et al. ChEBI in 2016: improved services and an expanding collection of metabolites. Nucleic Acids Res. 2016;44(D1):D1214–9.
Ammendola S, Lembo A, Battistoni A, Tagliatesta P, Ghisalberti C, Desideri A. 10-undecanhydroxamic acid, a hydroxamate derivative of the undecanoic acid, has strong antimicrobial activity through a mechanism that limits iron availability. FEMS Microbiol Lett. 2009;294(1):61–7.
Kabara JJ, Vrable R. Antimicrobial lipids: natural and synthetic fatty acids and monoglycerides. Lipids. 1977;12(9):753–9.
Avrahami D, Shai Y. Conjugation of a magainin analogue with lipophilic acids controls hydrophobicity, solution assembly, and cell selectivity. Biochemistry. 2002;41(7):2254–63.
Jones AM, Klun JA, Cantrell CL, Ragone D, Chauhan KR, Brown PN, et al. Isolation and identification of mosquito (Aedes aegypti ) biting deterrent fatty acids from male inflorescences of breadfruit (Artocarpus altilis (Parkinson) Fosberg). J Agric Food Chem. 2012;60(15):3867–73.
Ali A, Cantrell CL, Bernier UR, Duke SO, Schneider JC, Agramonte NM, et al. Aedes aegypti (Diptera: Culicidae) biting deterrence: structure–activity relationship of saturated and unsaturated fatty acids. J Med Entomol. 2012;49(6):1370–8.
Cruz-Estrada A, Ruiz-Sanchez E, Cristobal-Alejo J, Gonzalez-Coloma A, Andres MF, Gamboa-Angulo M. Medium-chain fatty acids from Eugenia winzerlingii leaves causing insect settling deterrent, nematicidal, and phytotoxic effects. Molecules. 2019;24(9):1724.
Brito-Madurro AG, Cuadros-Orellana S, Madurro JM, Martinez-Rossi N, Rossi A. Effect of undecanoic acid on the production of esterases and lipases by Aspergillus nidulans. Ann Microbiol. 2005;55(4):291–4.
Brito-Madurro AG, Prade RA, Madurro JM, Santos MA, Peres NT, Cursino-Santos JR, et al. A single amino acid substitution in one of the lipases of Aspergillus nidulans confers resistance to the antimycotic drug undecanoic acid. Biochem Genet. 2008;46(9–10):557–65.
Gershon H, Shanks L. Antifungal activity of fatty acids and derivatives: structure–activity relationships. In: Kabarra JJ, editor. The pharmacological effect of lipids. American Oil Chemists Society; 1978.
McDonough V, Stukey J, Cavanagh T. Mutations in erg4 affect the sensitivity of Saccharomyces cerevisiae to medium-chain fatty acids. Biochim Biophys Acta Mol Cell Biol Lipids. 2002;1581(3):109–18.
Shi D, Zhao Y, Yan H, Fu H, Shen Y, Lu G, et al. Antifungal effects of undecylenic acid on the biofilm formation of Candida albicans. Int J Clin Pharmacol Ther. 2016;54(5):343–53.
Mionic Ebersold M, Petrovic M, Fong WK, Bonvin D, Hofmann H, Milosevic I. Hexosomes with undecylenic acid efficient against Candida albicans. Nanomaterials (Basel). 2018;8(2):91.
Salini R, Sindhulakshmi M, Poongothai T, Pandian SK. Inhibition of quorum sensing mediated biofilm development and virulence in uropathogens by Hyptis suaveolens. Antonie Van Leeuwenhoek. 2015;107(4):1095–106.
Hortmann L, Rehm HJ. Inhibitory effect of undecanoic acid on the biosynthesis of long-chain fatty acids in Mortierella isabellina. Appl Microbiol Biotechnol. 1984;20(2):139–45.
Reverberi M, Punelli M, Smith CA, Zjalic S, Scarpari M, Scala V, et al. How peroxisomes affect aflatoxin biosynthesis in Aspergillus flavus. PLoS ONE. 2012;7(10):e48097.
Brito-Madurro AG, Cuadros-Orellana S, Martinez-Rossi NM, Rossi A. Undecanoic acid resistance in filamentous fungi: Identification and linkage mapping of the Aspergillus nidulans udaA gene. J Gen Appl Microbiol. 2005;51(1):47–9.
Persinoti GF, Martinez DA, Li W, Dogen A, Billmyre RB, Averette A, et al. Whole-genome analysis illustrates global clonal population structure of the ubiquitous dermatophyte pathogen Trichophyton rubrum. Genetics. 2018;208(4):1657–69.
Chaturvedi V, de Hoog GS. Onygenalean fungi as major human and animal pathogens. Mycopathologia. 2020;185(1):1–8.
Paião FG, Segato F, Cursino-Santos JR, Peres NT, Martinez-Rossi NM. Analysis of Trichophyton rubrum gene expression in response to cytotoxic drugs. FEMS Microbiol Lett. 2007;271(2):180–6.
Das SK, Banerjee AB. Effect of undecanoic acid on cell-permeability and respiration of Trichophyton rubrum. Acta Microbiol Pol. 1981;30(3):295–8.
Hiltunen JK, Mursula AM, Rottensteiner H, Wierenga RK, Kastaniotis AJ, Gurvitz A. The biochemistry of peroxisomal beta-oxidation in the yeast Saccharomyces cerevisiae. FEMS Microbiol Rev. 2003;27(1):35–64.
Shelest E. Transcription factors in fungi. FEMS Microbiol Lett. 2008;286(2):145–51.
Bahn YS. Exploiting fungal virulence-regulating transcription factors as novel antifungal drug targets. PLoS Pathog. 2015;11(7):e1004936.
Li XC, Jacob MR, Khan SI, Ashfaq MK, Babu KS, Agarwal AK, et al. Potent in vitro antifungal activities of naturally occurring acetylenic acids. Antimicrob Agents Chemother. 2008;52(7):2442–8.
Persinoti GF, Peres NTA, Jacob TR, Rossi A, Vencio RZ, Martinez-Rossi NM. RNA-sequencing analysis of Trichophyton rubrum transcriptome in response to sublethal doses of acriflavine. BMC Genomics. 2014;15(Suppl 7):S1.
Prasad R, Rawal MK. Efflux pump proteins in antifungal resistance. Front Pharmacol. 2014;5:202.
Aneke CI, Rhimi W, Otranto D, Cafarchia C. Synergistic effects of efflux pump modulators on the azole antifungal susceptibility of Microsporum canis. Mycopathologia. 2020;185(2):279–88.
Martins MP, Rossi A, Sanches PR, Martinez-Rossi NM. Differential expression of multidrug-resistance genes in Trichophyton rubrum. J Integr OMICS. 2019;9(2):1–81.
Fachin AL, Ferreira-Nozawa MS, Maccheroni W Jr, Martinez-Rossi NM. Role of the ABC transporter TruMDR2 in terbinafine, 4-nitroquinoline N-oxide and ethidium bromide susceptibility in Trichophyton rubrum. J Med Microbiol. 2006;55(Pt 8):1093–9.
Martins MP, Franceschini AC, Jacob TR, Rossi A, Martinez-Rossi NM. Compensatory expression of multidrug-resistance genes encoding ABC transporters in dermatophytes. J Med Microbiol. 2016;65(7):605–10.
Chandra J, Mukherjee PK, Ghannoum MA. Candida biofilms associated with CVC and medical devices. Mycoses. 2012;55:46–57.
Nett JE, Zarnowski R, Cabezas-Olcoz J, Brooks EG, Bernhardt J, Marchillo K, et al. Host contributions to construction of three device-associated Candida albicans biofilms. Infect Immun. 2015;83(12):4630–8.
Mayer FL, Wilson D, Hube B. Candida albicans pathogenicity mechanisms. Virulence. 2013;4(2):119–28.
Ma QX, Ola M, Iracane E, Butler G. Susceptibility to medium-chain fatty acids is associated with trisomy of chromosome 7 in Candida albicans. mSphere. 2019;4(3):e00402–19.
Selmecki A, Forche A, Berman J. Aneuploidy and isochromosome formation in drug-resistant Candida albicans. Science. 2006;313(5785):367–70.
Pavelka N, Rancati G, Zhu J, Bradford WD, Saraf A, Florens L, et al. Aneuploidy confers quantitative proteome changes and phenotypic variation in budding yeast. Nature. 2010;468(7321):321–5.
Cowen LE, Carpenter AE, Matangkasombut O, Fink GR, Lindquist S. Genetic architecture of Hsp90-dependent drug resistance. Eukaryot Cell. 2006;5(12):2184–8.
Cowen LE, Lindquist S. Hsp90 potentiates the rapid evolution of new traits: drug resistance in diverse fungi. Science. 2005;309(5744):2185–9.
Decaestecker E, Gaba S, Raeymaekers JAM, Stoks R, Van Kerckhoven L, Ebert D, et al. Host-parasite “Red Queen” dynamics archived in pond sediment. Nature. 2007;450(7171):870-U16.
Martinez-Rossi NM, Peres NT, Rossi A. Antifungal resistance mechanisms in dermatophytes. Mycopathologia. 2008;166(5–6):369–83.
Kelemen O, Convertini P, Zhang Z, Wen Y, Shen M, Falaleeva M, et al. Function of alternative splicing. Gene. 2013;514(1):1–30.
Kempken F. Alternative splicing in ascomycetes. Appl Microbiol Biotechnol. 2013;97(10):4235–41.
Mendes NS, Silva PM, Silva-Rocha R, Martinez-Rossi NM, Rossi A. Pre-mRNA splicing is modulated by antifungal drugs in the filamentous fungus Neurospora crassa. FEBS Open Bio. 2016;6(4):358–68.
Laloum T, Martin G, Duque P. Alternative splicing control of abiotic stress responses. Trends Plant Sci. 2018;23(2):140–50.
Chen W, Moore MJ. Spliceosomes. Curr Biol. 2015;25(5):R181-3.
Koncz C, Dejong F, Villacorta N, Szakonyi D, Koncz Z. The spliceosome-activating complex: molecular mechanisms underlying the function of a pleiotropic regulator. Front Plant Sci. 2012;3:9.
Bitencourt TA, Oliveira FB, Sanches PR, Rossi A, Martinez-Rossi NM. The prp4 kinase gene and related spliceosome factor genes in Trichophyton rubrum respond to nutrients and antifungals. J Med Microbiol. 2019;68(4):591–9.
Black DL. Mechanisms of alternative pre-messenger RNA splicing. Annu Rev Biochem. 2003;72:291–336.
Grutzmann K, Szafranski K, Pohl M, Voigt K, Petzold A, Schuster S. Fungal alternative splicing is associated with multicellular complexity and virulence: a genome-wide multi-species study. DNA Res. 2014;21(1):27–39.
Leal J, Squina FM, Freitas JS, Silva EM, Ono CJ, Martinez-Rossi NM, et al. A splice variant of the Neurospora crassa hex-1 transcript, which encodes the major protein of the Woronin body, is modulated by extracellular phosphate and pH changes. FEBS Lett. 2009;583(1):180–4.
Sieber P, Voigt K, Kammer P, Brunke S, Schuster S, Linde J. Comparative study on alternative splicing in human fungal pathogens suggests its involvement during host invasion. Front Microbiol. 2018;9:2313.
Roberts A, Pimentel H, Trapnell C, Pachter L. Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics. 2011;27(17):2325–9.
Goldstein LD, Cao Y, Pau G, Lawrence M, Wu TD, Seshagiri S, et al. prediction and quantification of splice events from RNA-Seq data. PLoS ONE. 2016;11(5):e0156132.
Hug N, Longman D, Caceres JF. Mechanism and regulation of the nonsense-mediated decay pathway. Nucleic Acids Res. 2016;44(4):1483–95.
Yu CH, Dang Y, Zhou Z, Wu C, Zhao F, Sachs MS, et al. Codon usage influences the local rate of translation elongation to regulate co-translational protein folding. Mol Cell. 2016;59(5):744–54.
Zhou Z, Dang Y, Zhou M, Yuan H, Liu Y. Codon usage biases co-evolve with transcription termination machinery to suppress premature cleavage and polyadenylation. Elife. 2018;7:29.
Funding
We thank the Brazilian funding agencies for the continuous support to our projects: São Paulo Research Foundation - FAPESP (proc. Nos. 2019/22596-9, 2018/11319-1, 2018/15458-6, 2015/23435-8, 2009/08411-4); National Council for Scientific and Technological Development -CNPq (proc. Nos. 305797/2017-4, and 304989/2017-7); Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - CAPES (Finance Code 001), and Fundação de Apoio ao Ensino, Pesquisa e Assistência - FAEPA.
Author information
Authors and Affiliations
Contributions
All authors wrote sections of the manuscript. NM-R, MM, and AR edited the manuscript. All authors read and approved the submitted version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Research involving human participants and/or animals
This manuscript does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: Vishnu Chaturvedi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rossi, A., Martins, M.P., Bitencourt, T.A. et al. Reassessing the Use of Undecanoic Acid as a Therapeutic Strategy for Treating Fungal Infections. Mycopathologia 186, 327–340 (2021). https://doi.org/10.1007/s11046-021-00550-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11046-021-00550-4